Nucleus · Proton-Electron model
RealQM applied to atomic nuclei: protons and electrons as charged density domains interacting via Coulomb forces. The old proton-electron model of the nucleus (4 protons + 2 electrons = He-4, total charge +2) revisited with modern computational methods.
RealNucleus — the same Coulomb framework, at femtometer scale
The math. Same multiphase Coulomb continuum-mechanics formulation used for atoms and molecules elsewhere in this gallery: each proton and each electron carries its own non-overlapping unit-charge density domain in 3D, with Bernoulli free boundaries between adjacent species, and energy minimisation over all densities and all boundaries. Equal mass mi=1 for both species in the model.
Mutual confinement, no strong force. Protons hold electrons in place (outer positive cage attracting the inner negative core) and electrons hold protons in place (inner negative anchor pulling the outer positive shell inward against its own repulsion). Neither species is confined by an external potential. Crucially, no strong nuclear force is invoked — pure Coulomb suffices.
Scale rescaling. Coulomb's 1/r is scale-invariant: the same variational solution at femtometer (instead of Bohr-radius) length scales multiplies energies by ~105. Net conversion: 1 Ha → 2.72 MeV. Geometry, boundary topology, and the variational minimum carry over unchanged; only units change.
Two implementations on the same model. • 3D polyhedral (light nuclei, Z ≤ ~8) — electrons at vertices of an inner Platonic solid, protons on a larger outer polytope (He-4, Li-6, Be-8, C-12, O-16, Mg-24, Ca-40). Reproduces nuclear-binding magnitudes to within ~25% across the small-Z range.
• 1D radial multi-shell (any Z) — spherically symmetric reduction: homogenised inner electron core + stack of concentric proton shells. Shell count serves as the calibration knob that fits the experimental E/A curve across the full periodic table, including the Fe-56 turnover — still pure Coulomb, no new physics.
The central claim. Both fusion-side and fission-side energetics live inside one variational Coulomb principle. The nuclear demonstration is not just a proof-of-concept analogy — it is a claim that the same framework that handles atoms, molecules, hydrogen bonding, and protein folding extends quantitatively to the nuclear scale without adding any new force law.
RealQM for the Atomic Nucleus (PDF) · Blog
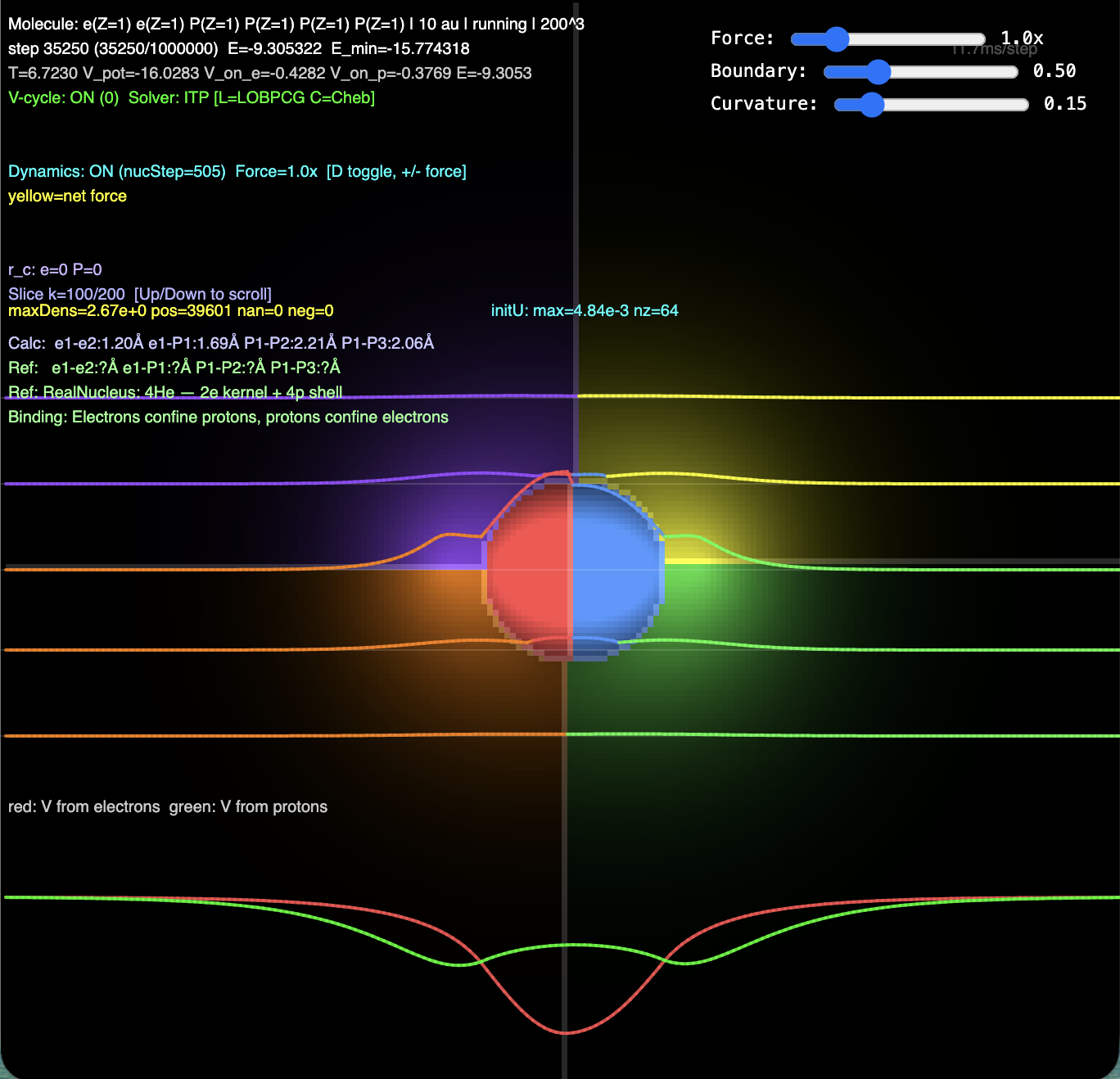
He-4 Nucleus (2e + 4p) — A Happy Marriage
Green potential well (from electrons) confines protons outward.
Neither species has an external confining potential — they hold each other in place through mutual Coulomb attraction. A happy marriage where electrons need protons and protons need electrons.
1 Electron + 4 Protons
R Sweep
2e + 1P (Z=4)
2e+1P R Sweep
4e + 8p
Li-6 (3e + 6p)
Be-8 (4e + 8p)
C-12 (6e + 12p)
O-16 (8e + 16p)
Mg-24 (12e + 24p)
Ca-40 (20e + 40p)
Packing Model with Continuous Core Expansion — full B/A curve
|E/A|(Z) = 1.36 / (2Re + 2γZ + c/Z) MeV.
Three parameters (c, Re, γ), single proton shell of 2Z protons
(size d=c/Z) around a continuously expanding electron core
(Reeff=Re+γZ). No piecewise structure, no
threshold, no Zbreak. The Fe-Ni binding maximum emerges as the saddle
point of the c/Z vs γZ competition in the denominator:
Zpeak = √(c/(2γ)) ≈ 22−29 for
c ≈ 0.5, γ ≈ 3−5×10−4. With these
values the model reproduces the full experimental B/A curve from H-2 to U-238
— rise to the Fe-Ni peak (8.79 MeV/A) and post-iron decline to U-238 (7.57
MeV/A) — in one analytic expression.The 3D polyhedral picture for Z ≤ 8 is an angularly-resolved alternative to the spherical reduction (electrons and protons at Platonic vertices: tetrahedron, cube, icosahedron). Same physics, different representation: angularly resolved vs angularly averaged.
Geometric essence: electrons are large, protons are small. Each electron's domain in the core: ~Re/Z1/3. Each proton: ~c/Z. Size ratio (electron/proton) ~ ReZ2/3/c. The Z2/3 scaling matches Weizsäcker's surface term, but emerges geometrically from the proton-electron size hierarchy rather than from a postulated surface penalty. The post-Fe-56 decline emerges from continuous core expansion — the natural geometric response when one shell cannot accommodate more constituents at the original packing.
Both Coulomb repulsions are present even though only one shows up in the formula. Proton-proton intra-shell repulsion is explicit (cancels with the leading $-2Z^2/r*$ proton-electron attraction, leaving the $-Z/r*$ residual); without it the model would predict $-Z^2/r*$, two orders of magnitude over-bound. Electron-electron repulsion is implicit: encoded both in the fitted Re (the value variational balance picks for the inner-core size at small Z) AND in the expansion rate γ (the rate at which Re must grow to keep e-e repulsion per nucleon manageable as N grows past ~28). The post-Fe-56 decline is the macroscopic shadow of e-e repulsion in the core. Full 3D polyhedral and radial multi-shell simulations include both repulsions explicitly and converge to negative energies (Li-6: −9.57, Be-8: −25.88, C-12: −25.90 Ha), confirming variational stability. The reduced formula doesn't contradict these — it summarises them.
Strong force not required anywhere in this chain. Same Coulomb-with-Bernoulli-boundary formulation that handles atoms, molecules, H-bonding, and protein folding accounts for the full nuclear B/A curve through one mechanism.
6-Shell Proton Model (1D prototype)
He-4 (point kernel)
Deuterium
D + T → He-4 + n — 3D fusion simulation
D + T → He-4 + n + 17.6 MeV
(the basis of ITER, NIF, and all proposed fusion reactors) modeled as a
multiphase 3D Coulomb continuum-mechanics simulation. In the proton-electron
picture (neutron = p + e), the constituents are:• D = deuterium (H-2) = 2 protons + 1 electron, net +1
• T = tritium (H-3) = 3 protons + 2 electrons, net +1
• He-4 = alpha particle = 4 protons + 2 electrons, net +2
• n = neutron = 1 proton + 1 electron, net 0
Total: 5 protons + 3 electrons on both sides — the reaction is a pure topological rearrangement of the same 8 constituents from two clusters (D + T) into one tight cluster (He-4) plus a free pair (n) that drifts off.
Result. Starting with the 8 subdomains in a D-blob + T-blob configuration with all boundaries free and dynamics on, the variational principle spontaneously finds the He-4 + n configuration. Total energy descends from ∼−6 Ha (D + T separated) to ∼−11.5 Ha (He-4 + ejected neutron), ΔE ≈ 5.5 Ha drop. Rescaled at nuclear scale (1 Ha → 2.72 MeV): ΔE ≈ 15 MeV released vs experimental Q-value 17.6 MeV — 85% agreement from pure Coulomb with no strong-force input, no reaction rules, no transition-state ansatz.
Set Rsep via URL:
?rsep=0.5
(close, fast merge), ?rsep=1.5 (moderate), ?rsep=2.5
(further, needs to overcome barrier).